Research of Yokoyama Lab
(Last update 2023.4)
-- Structural biology of membrane signalling proteins or drug-target proteins
using X-ray crystallography --
I try to determine three-dimensional structures of membrane signalling proteins or drug-target proteins.
Under techniques such as X-ray crystallography and molecular biology, proteins are expressed
using E. coli or insect cells, purified, and crystallized.
And then protein structures are determined.
I want to elucidate structures and functions of proteins.
Now I proceed the following researches.
1. Structural and functional elucidation of membrane signalling proteins
・Stomatin, a lipid raft protein
・Podocin, a causative protein of nephrotic syndrome
2. Elucidation of binding mechanism of protein-inhibitor complex to develop anti-cancer drugs
・Kinesin CENP-E - inhibitor complex (Collaborative research with Prof. Asai and Associate Prof. Sawada, Center for Drug Discovery,
Univ. of Shizuoka)
(Acta Crystallog. D 2021,
Press Release230403)
・Claudin, a tight junction protein - inhibitor complex (Collaborative research with Prof. Ikari, Gifu Pharm. Univ.)
・Signaling lymphocytic activation molecule family 7 (SLAMF7) - antibody complex
Protein Data Bank (PDB) ID:
6M4I
(On-campus collaborative research)
・Ir complex - calmodulin(Collaborative research with Aoki Lab.)
・Matrix metalloproteinase 21 (MMP21) (Collaborative research with Kon Lab., Gra. Sch of Biol. Sci.)
・Glyoxalase I (GLO I) (Collaborative research with Takasawa Lab.)
・Poly(ADP-ribose) glycohydrolase (PARG) (Collaborative research with Tanuma Lab., Res. Inst. Sci. Tech.)
・Mitochondrial intermediate peptidase (MIPEP) (Collaborative research with Higami Lab.)
・Phosphoinositide 3-kinase (PI3K) - caffeine complex、
Poly(ADP-ribose) polymerase-1 (PARP1) - caffeine metabolite complex (Collaborative research with Aoyama Lab, Dr. Kawano)
I have done the following researches.
1. Structural and functional elucidation of membrane signalling proteins
(1) Elucidation of regulation mechanism of lipid raft protein stomatin and its specific membrane-bound protease (Now in progress)
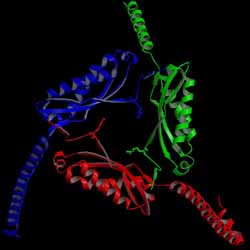 Stomatin is a major integral membrane protein of human erythrocytes,
the absence of which is associated with a form of hemolytic anemia known as hereditary stomatocytosis.
The function of stomatin is not fully understood.
As a lipid raft protein, it could act as an oligomeric scaffolding protein or
as an active signaling component involved in vesicle trafficking.
Stomatin is a major integral membrane protein of human erythrocytes,
the absence of which is associated with a form of hemolytic anemia known as hereditary stomatocytosis.
The function of stomatin is not fully understood.
As a lipid raft protein, it could act as an oligomeric scaffolding protein or
as an active signaling component involved in vesicle trafficking.
In archaeal and bacterial species, p-stomatin (prokaryotic stomatin) and
STOPP (STomatin Operon Partner Protein) genes probably form an operon.
We determined the first crystal structure of the core domain of p-stomatin.
It forms a novel homotrimeric structure (left figure).
(J. Mol. Biol. 2008,
Yakugaku Zasshi 2010)
While, mouse stomatin forms a homodimer. We elucidated the region to determine whether stomatin domains form dimers or trimers.
(Biochem. Biophys. Rep. 2022)
N-terminal region of STOPP (STOPP-N) is a serine protease,
and specifically cleaves p-stomatin.
We determined crystal structures of STOPP-N and its complex with substrate peptide.
STOPP-N forms a homodimer.
A structural comparison revealed binding of the substrate to cause a large rotational and translational displacement
between protomers.
(J. Biol. Chem. 2005,
J. Mol. Biol. 2006,
J. Synchrotron. Rad. 2008,
Biochemistry 2012,
J. Synchrotron. Rad. 2013,
Acta Crystallog. D 2020)
C-terminal region of STOPP (STOPP-C) has an OB(Oligosaccharide/oligonucleotide Binding)-fold domain,
and assembles into oligomers.
STOPP-C may function as a scaffold protein to form multimeric assembly of STOPP and stomatin.
(Biochimie 2013,
FEBS Open Bio 2014)
(These are collaborative researches with Dr. Ikuo Matsui who belongs to
National Institute of Advanced Industrial Science and Technology (AIST).)
PDB ID:
3BK6
2DEO
3BPP
3VIV
3WG5
3WWV
6M4B
8GN9
(2) Elucidation of peptide-binding and transporting mechanism of oligopeptide permease OppA
Bacterial oligopeptide permease is involved in signal transduction by binding and transporting of peptides.
Some archaea produce membrane vesicles, and their major protein is OppA.
In order to elucidate peptide-binding and transporting mechanism in archaea,
we tried to determine structures of OppA and its complexes with peptides.
(Crystallogr. Rep. 2021)
2. Drug-target proteins: elucidation of allosteric inhibition mechanism in kinesin spindle protein Eg5
Kinesin spindle protein Eg5 is a mitotic spindle motor protein with ATPase activity.
Inhibition of Eg5 causes cell cycle arrest in mitosis and subsequent apoptotic cell death.
Human Eg5 is an attractive target for novel anticancer therapies.
We determined the structure of Eg5 motor domain in complex with its biphenyl-type ATP-competitive inhibitor.
This structure will provide useful information for development of novel types of allosteric drugs.
(ACS Chem. Biol. 2015)
We also determined the structures of Eg5 motor domain in complex with STLC-type ATP-uncompetitive inhibors, PVEI0021 or PVEI0138.
(ACS Omega 2018)
(These are collaborative researches with Prof. Asai and Associate Prof. Sawada, Center for Drug Discovery, Univ. of Shizuoka.)
PDB ID:
3WPN
5ZO7
5ZO8
5ZO9
3. Elucidation of metal-ion uptake mechanism of Helicobacter pylori
neutrophil-activating protein
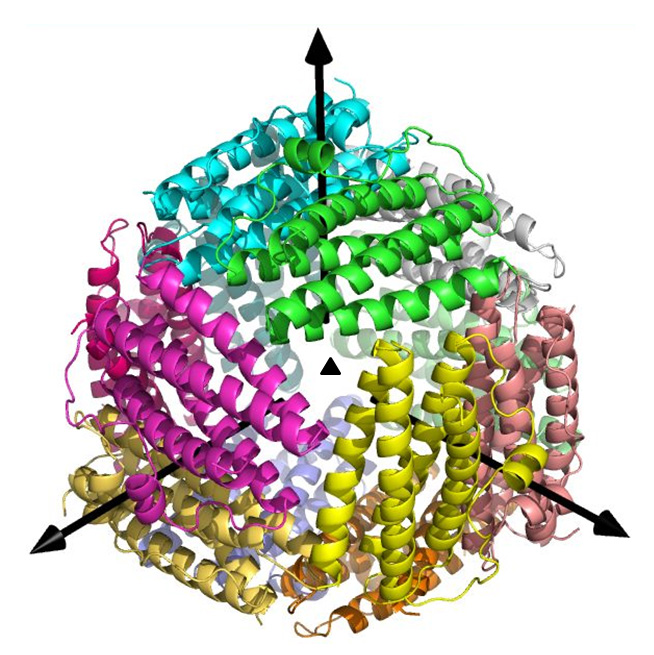 Helicobacter pylori causes severe diseases such as chronic gastritis, peptic ulcers,
and stomach cancers.
H. pylori neutrophil-activating protein (HP-NAP) attracts and activates neutrophils,
and induces the production of reactive oxygen radicals.
HP-NAP is a Dps-like iron storage protein forming a dodecameric shell.
We determined crystal structures of Fe, Zn, or Cd ion-bound form or apo form of HP-NAP.
(Acta Crystallog. F 2012,
Biochem. Biophys. Res. Commun. 2012,
Biomolecules 2014)
Helicobacter pylori causes severe diseases such as chronic gastritis, peptic ulcers,
and stomach cancers.
H. pylori neutrophil-activating protein (HP-NAP) attracts and activates neutrophils,
and induces the production of reactive oxygen radicals.
HP-NAP is a Dps-like iron storage protein forming a dodecameric shell.
We determined crystal structures of Fe, Zn, or Cd ion-bound form or apo form of HP-NAP.
(Acta Crystallog. F 2012,
Biochem. Biophys. Res. Commun. 2012,
Biomolecules 2014)
Iron loading causes a series of conformational changes of some amino acid residues
at the ferroxidase center.
Metal-ions are found inside of the negatively-charged pore, and then
the pore is suitable for metal ions to pass through.
PDB ID:
3T9J
3TA8
4EVB
4EVC
4EVD
4EVE
4. Structural elucidation of UV-damaged DNA (6-4) photoproduct and its specific antibody Fab
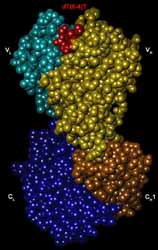 DNA photoproducts produced by ultraviolet irradiation cause mutations,
cellular transformation, and cell death.
Pyrimidine (6-4) pyrimidone DNA photoproducts are more mutagenic than cyclobutane pyrimidine dimers.
DNA photoproducts produced by ultraviolet irradiation cause mutations,
cellular transformation, and cell death.
Pyrimidine (6-4) pyrimidone DNA photoproducts are more mutagenic than cyclobutane pyrimidine dimers.
We determined the structures of (6-4) photoproduct in complex with its antibody Fab.
5'-thymine and 3'-pyrimidone bases of dT(6-4)T are in half-chair and planar conformations, respectively,
and are nearly perpendicular to each other.
dT(6-4)T is fully accommodated in concaved pocket of Fab.
(J. Mol. Biol. 2000,
Acta Crystallog. D 2012,
Acta Crystallog. D 2013,
Int. J. Mol. Sci. 2014,
Acta Crystallog. F 2019,
Sci. Rep. 2019)
PDB ID:
1EHL
1KEG
3VW3
6IDG
6IDH
6KDH
6KDI
 Stomatin is a major integral membrane protein of human erythrocytes,
the absence of which is associated with a form of hemolytic anemia known as hereditary stomatocytosis.
The function of stomatin is not fully understood.
As a lipid raft protein, it could act as an oligomeric scaffolding protein or
as an active signaling component involved in vesicle trafficking.
Stomatin is a major integral membrane protein of human erythrocytes,
the absence of which is associated with a form of hemolytic anemia known as hereditary stomatocytosis.
The function of stomatin is not fully understood.
As a lipid raft protein, it could act as an oligomeric scaffolding protein or
as an active signaling component involved in vesicle trafficking. Helicobacter pylori causes severe diseases such as chronic gastritis, peptic ulcers,
and stomach cancers.
H. pylori neutrophil-activating protein (HP-NAP) attracts and activates neutrophils,
and induces the production of reactive oxygen radicals.
HP-NAP is a Dps-like iron storage protein forming a dodecameric shell.
We determined crystal structures of Fe, Zn, or Cd ion-bound form or apo form of HP-NAP.
(Acta Crystallog. F 2012,
Biochem. Biophys. Res. Commun. 2012,
Biomolecules 2014)
Helicobacter pylori causes severe diseases such as chronic gastritis, peptic ulcers,
and stomach cancers.
H. pylori neutrophil-activating protein (HP-NAP) attracts and activates neutrophils,
and induces the production of reactive oxygen radicals.
HP-NAP is a Dps-like iron storage protein forming a dodecameric shell.
We determined crystal structures of Fe, Zn, or Cd ion-bound form or apo form of HP-NAP.
(Acta Crystallog. F 2012,
Biochem. Biophys. Res. Commun. 2012,
Biomolecules 2014) DNA photoproducts produced by ultraviolet irradiation cause mutations,
cellular transformation, and cell death.
Pyrimidine (6-4) pyrimidone DNA photoproducts are more mutagenic than cyclobutane pyrimidine dimers.
DNA photoproducts produced by ultraviolet irradiation cause mutations,
cellular transformation, and cell death.
Pyrimidine (6-4) pyrimidone DNA photoproducts are more mutagenic than cyclobutane pyrimidine dimers.